Structure of Family 5 Uracil-DNA Glycosylase Bound to DNA Reveals Insights into the Mechanism for Substrate Recognition and Catalysis
Kosaka, H., Nakagawa, N., Masui, R., Kuramitsu, S.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| uracil-DNA glycosylase | C [auth A] | 219 | Thermus thermophilus HB8 | Mutation(s): 0 Gene Names: ttUDGB EC: 3.2.2 | 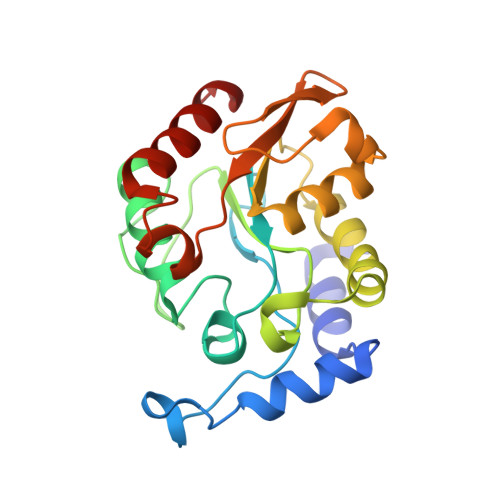 |
UniProt | |||||
Find proteins for Q5SJ65 (Thermus thermophilus (strain ATCC 27634 / DSM 579 / HB8)) Explore Q5SJ65 Go to UniProtKB: Q5SJ65 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q5SJ65 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Find similar nucleic acids by: Sequence | 3D Structure
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | Organism | Image | |
| 5'-D(*AP*TP*GP*TP*TP*GP*CP*(D1P)P*TP*TP*AP*GP*TP*CP*C)-3' | A [auth C] | 15 | N/A | 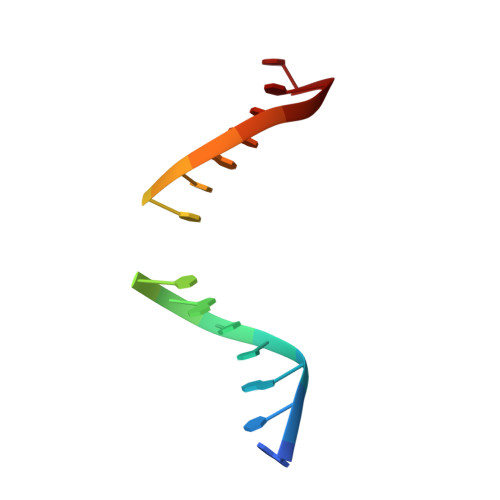 | |
Sequence AnnotationsExpand | |||||
| |||||
Find similar nucleic acids by: Sequence | 3D Structure
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | Organism | Image | |
| 5'-D(*GP*GP*AP*CP*TP*AP*AP*CP*GP*CP*AP*AP*CP*A)-3' | B [auth D] | 14 | N/A | 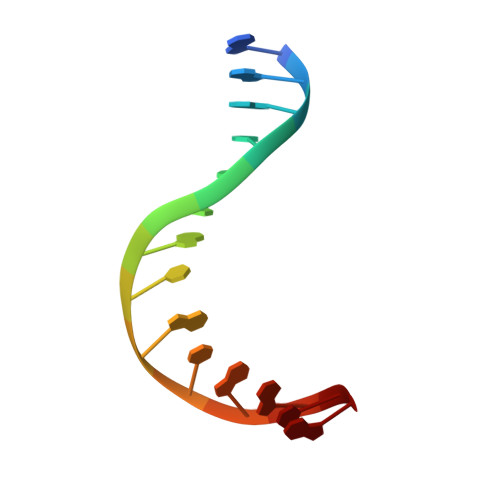 | |
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 4 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| SF4 Query on SF4 | E [auth A] | IRON/SULFUR CLUSTER Fe4 S4 LJBDFODJNLIPKO-UHFFFAOYSA-N |  | ||
| 2HP Query on 2HP | D, F [auth A] | DIHYDROGENPHOSPHATE ION H2 O4 P NBIIXXVUZAFLBC-UHFFFAOYSA-M |  | ||
| GOL Query on GOL | G [auth A], H [auth A] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| EOH Query on EOH | I [auth A] | ETHANOL C2 H6 O LFQSCWFLJHTTHZ-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 68.868 | α = 90 |
| b = 148.754 | β = 90 |
| c = 93.086 | γ = 90 |
| Software Name | Purpose |
|---|---|
| ADSC | data collection |
| HKL-2000 | data reduction |
| MLPHARE | phasing |
| CNS | refinement |
| HKL-2000 | data scaling |